Know your (true) correlation in life sciences data
20 January 2020
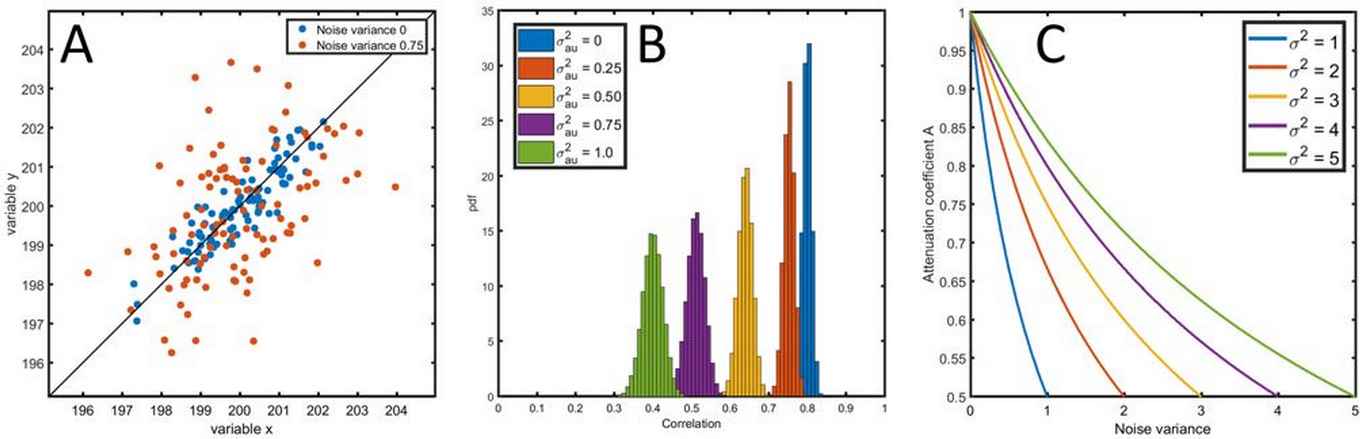
The use of correlation coefficients can be limited to simple experimental analysis, but they are also basic ingredients for sophisticated multivariate (or “big”) data analysis methods. It is therefore important to have reliable estimates for correlation coefficients. In current-day life sciences, comprehensive measurement techniques are used to measure all kinds of complex data, such as metabolites, proteins, gene-expressions and more. All these measurement techniques have errors. Whereas in the old days, with simple measurements, the errors were also simple, that is not the case anymore. Errors are heterogeneous, non-constant and not independent. This hampers the quality of the estimated correlation coefficients seriously and it can even change the sign of the ‘true’ biological correlation. If this is the case, it can severely affect the outcome of an experiment.
In their findings, the researchers discuss the different types of errors as present in modern comprehensive life science data and show with theory, simulations and real-life data how these affect the correlation coefficients. They also opt ways to improve the estimation of such coefficients. Based on their analysis, the researchers warn that in general correlation coefficients are trusted too much on face value and they hope to have triggered some doubts and pointed to precautions regarding the use of correlations for future experiments.